I am a computational biologist and group leader at Illumina's Artificial Intelligence Laboratory, where my research is focused on understanding the functional impact of human genetic variation. I am also a member of the Genotype-Tissue Expression (GTEx) Project Consortium, which built an atlas of genetic effects on gene expression across human tissues, and the recently launched Developmental GTEx (dGTEx) project. Previously, I was a group leader at the Broad Institute, within the Getz lab.
Selected papers
H. Xu, L. Chen, M. Sun, K. Jean-Baptiste, D. Cao, X. Zhou, S. Wong, C. Xiao, T. Liu, V. Quijano, N. Brandes, J. Germino, V. Ntranos, C.J. Ye, F. Aguet*, K. Farh*, "Single cell sequencing as a general variant interpretation assay", bioRxiv, 2023.
 G. Eraslan, E. Drokhlyansky, S. Anand, E. Fiskin, A. Subramanian, M. Slyper, J. Wang, N. Van Wittenberghe, J.M. Rouhana, J. Waldman, O. Ashenberg, M. Lek, D. Dionne, T.S. Win, M.S. Cuoco, O. Kuksenko, A.M. Tsankov, P.A. Branton, J.L. Marshall, A. Greka, G. Getz, A.V. Segrè*, F. Aguet*, O. Rozenblatt-Rosen*, K.G. Ardlie*, A. Regev*, "Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function," Science 376, eabl4290, 2022. Coverage: [Science Perspective] [Broad Institute]
G. Eraslan, E. Drokhlyansky, S. Anand, E. Fiskin, A. Subramanian, M. Slyper, J. Wang, N. Van Wittenberghe, J.M. Rouhana, J. Waldman, O. Ashenberg, M. Lek, D. Dionne, T.S. Win, M.S. Cuoco, O. Kuksenko, A.M. Tsankov, P.A. Branton, J.L. Marshall, A. Greka, G. Getz, A.V. Segrè*, F. Aguet*, O. Rozenblatt-Rosen*, K.G. Ardlie*, A. Regev*, "Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function," Science 376, eabl4290, 2022. Coverage: [Science Perspective] [Broad Institute]
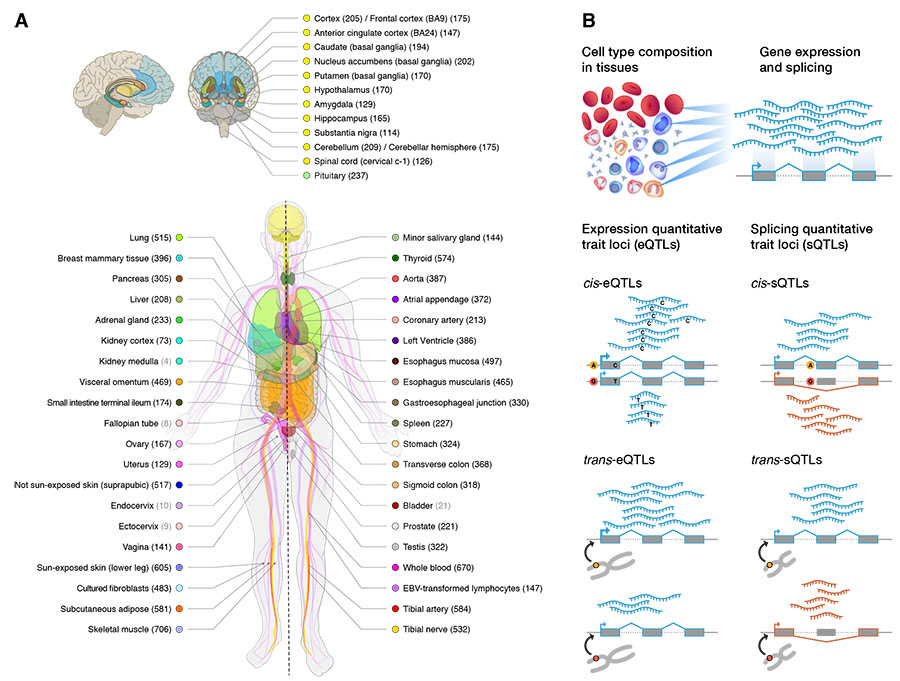
 GTEx Consortium (first & co-corresponding author), "The GTEx Consortium atlas of genetic regulatory effects across human tissues," Science 369, 1318-1330, 2020. Coverage: [Science News] [Broad Institute] [NYGC]
GTEx Consortium (first & co-corresponding author), "The GTEx Consortium atlas of genetic regulatory effects across human tissues," Science 369, 1318-1330, 2020. Coverage: [Science News] [Broad Institute] [NYGC]
S. Kim-Hellmuth*, F. Aguet*, M. Oliva, M. Muñoz-Aguirre, S. Kasela, V. Wucher, S.E. Castel, A.R. Hamel, A. Viñuela, A.L. Roberts, S. Mangul, X. Wen, G. Wang, A.N. Barbeira, D. Garrido-Martín, B. Nadel, Y. Zou, R. Bonazzola, J. Quan, A. Brown, A. Martinez-Perez, J.M. Soria, GTEx Consortium, G. Getz, E.T. Dermitzakis, K.S. Small, M. Stephens, H.S. Xi, H.K. Im, R. Guigó, A.V. Segrè, B.E. Stranger, K.G. Ardlie, T. Lappalainen "Cell type specific genetic regulation of gene expression across human tissues," Science 369, eaaz8528, 2020.