I am a computational biologist and currently lead Research and Computational Biology at Predicta Biosciences, a precision oncology company integrating cutting-edge multiomics and cell-based liquid biopsies to develop transformative diagnostic and therapeutic products. Previously, I was an Associate Director at Illumina's Artificial Intelligence Laboratory and a Group Leader at the Broad Institute, where my research was focused on understanding the genetic basis of human diseases. I am/was a member of several research consortia working towards this goal, including the Genotype-Tissue Expression (GTEx) Project Consortium, which built an atlas of genetic effects on gene expression across human tissues, the Trans-Omics for Precision Medicine (TOPMed) Program, the Clinical Proteomic Tumor Analysis Consortium (CPTAC), and the recently launched Developmental GTEx (dGTEx) project.
Selected publications
P. Orchard, ..., NHLBI TOPMed Consortium, F. Aguet*, L. Scott*, L.M. Raffield*, S.C.J. Parker*, Cross-cohort analysis of expression and splicing quantitative trait loci in TOPMed”, medRxiv, 2025.
H. Xu, L. Chen, M. Sun, K. Jean-Baptiste, D. Cao, X. Zhou, S. Wong, C. Xiao, T. Liu, V. Quijano, N. Brandes, J. Germino, V. Ntranos, C.J. Ye, F. Aguet*, K. Farh*, "Single cell sequencing as a general variant interpretation assay", bioRxiv, 2023.
 G. Eraslan, E. Drokhlyansky, S. Anand, E. Fiskin, A. Subramanian, M. Slyper, J. Wang, N. Van Wittenberghe, J.M. Rouhana, J. Waldman, O. Ashenberg, M. Lek, D. Dionne, T.S. Win, M.S. Cuoco, O. Kuksenko, A.M. Tsankov, P.A. Branton, J.L. Marshall, A. Greka, G. Getz, A.V. Segrè*, F. Aguet*, O. Rozenblatt-Rosen*, K.G. Ardlie*, A. Regev*, "Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function," Science 376, eabl4290, 2022. Coverage: [Science Perspective] [Broad Institute]
G. Eraslan, E. Drokhlyansky, S. Anand, E. Fiskin, A. Subramanian, M. Slyper, J. Wang, N. Van Wittenberghe, J.M. Rouhana, J. Waldman, O. Ashenberg, M. Lek, D. Dionne, T.S. Win, M.S. Cuoco, O. Kuksenko, A.M. Tsankov, P.A. Branton, J.L. Marshall, A. Greka, G. Getz, A.V. Segrè*, F. Aguet*, O. Rozenblatt-Rosen*, K.G. Ardlie*, A. Regev*, "Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function," Science 376, eabl4290, 2022. Coverage: [Science Perspective] [Broad Institute]
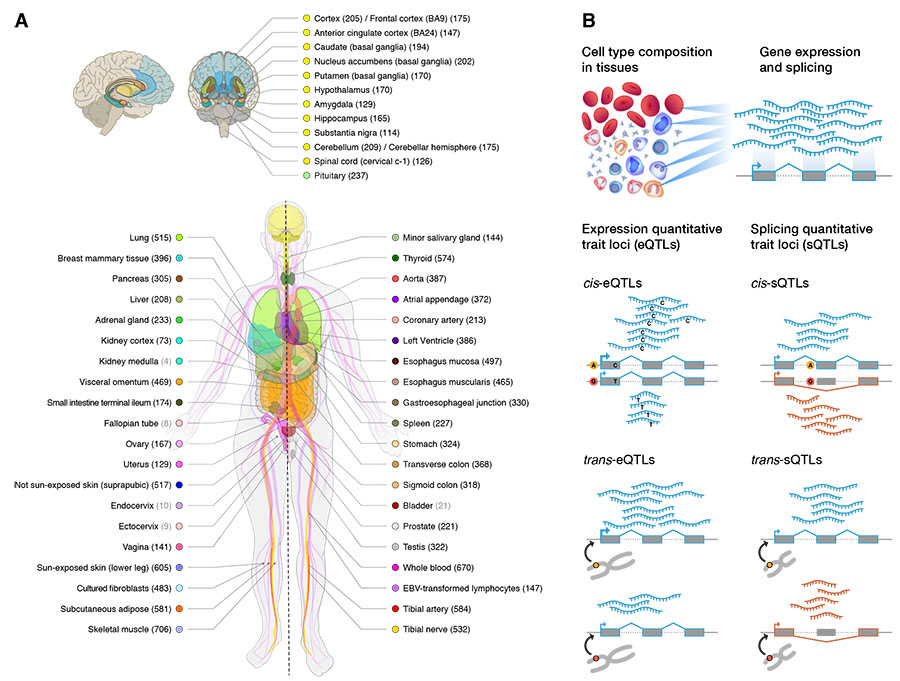
 GTEx Consortium (first & co-corresponding author), "The GTEx Consortium atlas of genetic regulatory effects across human tissues," Science 369, 1318-1330, 2020. Coverage: [Science News] [Broad Institute] [NYGC]
GTEx Consortium (first & co-corresponding author), "The GTEx Consortium atlas of genetic regulatory effects across human tissues," Science 369, 1318-1330, 2020. Coverage: [Science News] [Broad Institute] [NYGC]
S. Kim-Hellmuth*, F. Aguet*, M. Oliva, M. Muñoz-Aguirre, S. Kasela, V. Wucher, S.E. Castel, A.R. Hamel, A. Viñuela, A.L. Roberts, S. Mangul, X. Wen, G. Wang, A.N. Barbeira, D. Garrido-Martín, B. Nadel, Y. Zou, R. Bonazzola, J. Quan, A. Brown, A. Martinez-Perez, J.M. Soria, GTEx Consortium, G. Getz, E.T. Dermitzakis, K.S. Small, M. Stephens, H.S. Xi, H.K. Im, R. Guigó, A.V. Segrè, B.E. Stranger, K.G. Ardlie, T. Lappalainen "Cell type specific genetic regulation of gene expression across human tissues," Science 369, eaaz8528, 2020.